Snapping shrimp make plenty of noise, around 220 dB (play this video for a natural soundtrack to this post). They’ve also received a bunch of scientific attention. This is partly for their hard-to-miss chorus but it’s also because some species are super social, or in zoological jargon, eusocial.
Eusocial is considered the highest level of social organization and describes species that live in groups with overlapping generations (requirement #1), divide labor into reproductive and nonreproductive jobs (requirement #2), and participate in the cooperative care of juveniles (requirement #3). How this situation squares with natural selection has intrigued evolutionary biologists starting with Darwin himself and is wonderfully summarized in this Nature Education article.
Close to 50,000 species have been classified as eusocial. Most ants and termites are eusocial as are many bees and wasps. Some aphids, thrips, and beetles also show eusocial behavior as do certain naked mole rats. In addition several primate species have been classified as primitively eusocial.
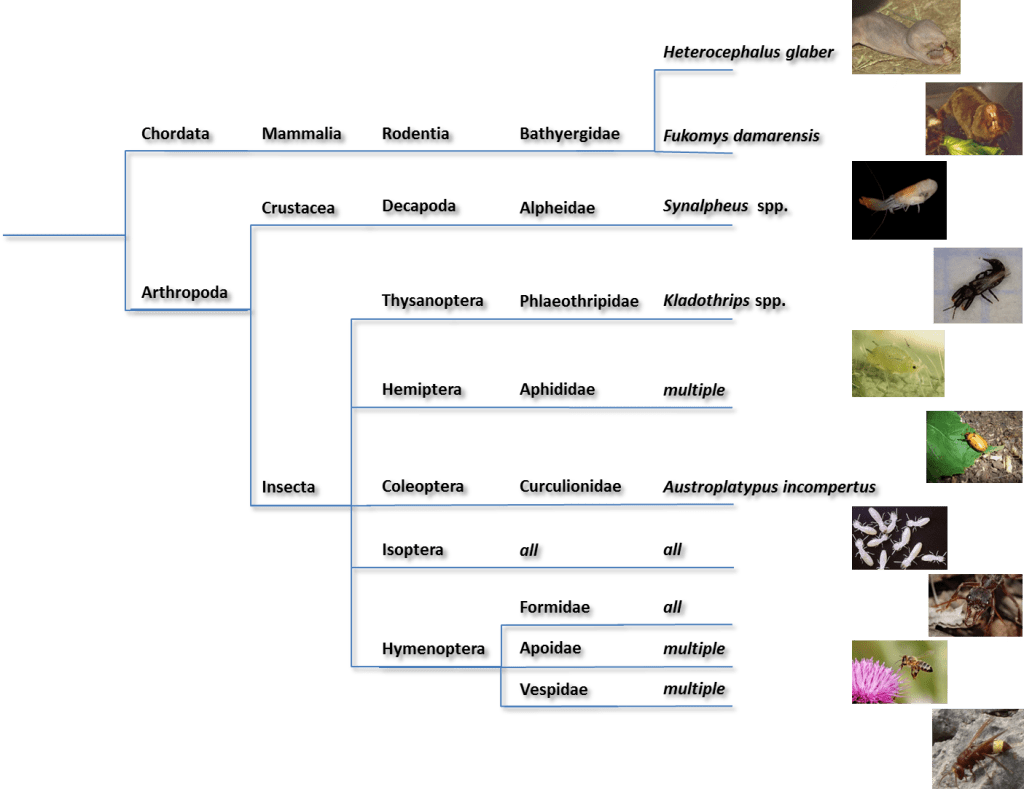
However, under the sea, it’s a different story.
Only eight marine animals are considered eusocial. They all belong to the Synalpheus genus of snapping shrimp. Their names are Synalpheus brooksi, Synalpheus chacei, Synalpheus elizabethae, Synalpheus filidigitus, Synalpheus rathbunae, Synalpheus regalis, Synalpheus microneptunus, and Synalpheus duffyi. But for convenience and mnemonics let’s call them the eusocial eight. The eusocial eight all live in colonies of 100-500 closely related individuals with one reproductive female and many male defenders. (Interesting aside: The latter have extra-large claws that produce high-speed waterjets capable of stunning other animals.)
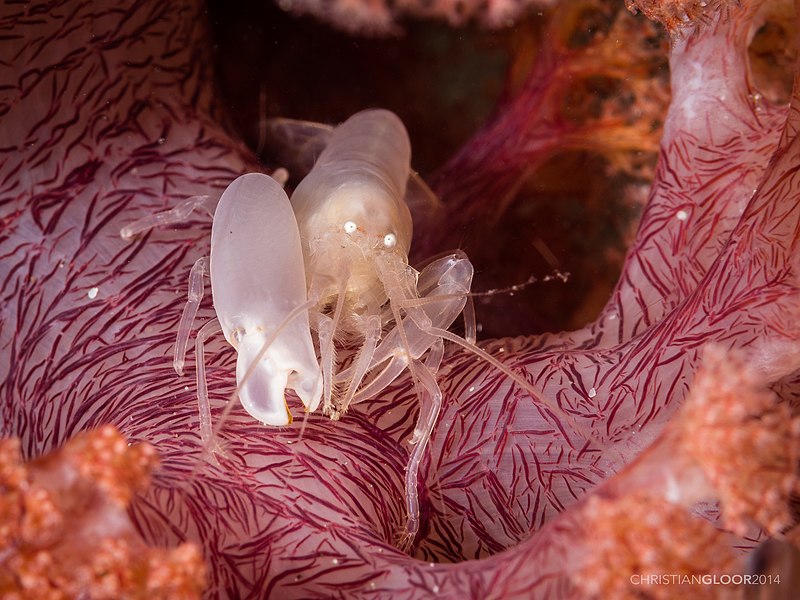

The eusocial eight can be found in tropical oceans where they have a highly dependent and highly specific relationship with certain sponge species which they depend on for both food and shelter. These sponges are in short supply and most are already inhabited (kind of like today’s housing market). As a result, members of these shrimp colonies have adopted a fortress defense strategy where individuals choose an extreme form of cooperation to defend a highly valuable resource. Ecological studies indicate that this strategy pays off – the eusocial eight are more abundant, occupy more sponges, and have broader host ranges than closely related but non-social shrimp species.
Phylogeny studies of the Synalpheus genus based on both genetic data and physical traits show an interesting pattern. Although eusociality appears only in this one genus among all marine animals, within the genus the trait pops up in different branches. This suggests that the trait has independently evolved multiple times.
This evolutionary pattern attracted the attention of scientists at Columbia University who study the causes and consequences of sociality as well as species responses to environmental uncertainty. They planned to reconstruct the evolution of social diversity within the Synalpehus genus using whole-genome sequencing. In the process, they discovered something completely unexpected. They found that genome sizes within the genus varied tremendously (between 4,700 Mbp and 20,300 Mbp) and that it was consistently the social shrimp that had the super-sized genomes. A deeper dive into the nucleotide data revealed the molecular biology behind this pattern – the eusocial eight’s genomes were all jam-packed with jumping genes.
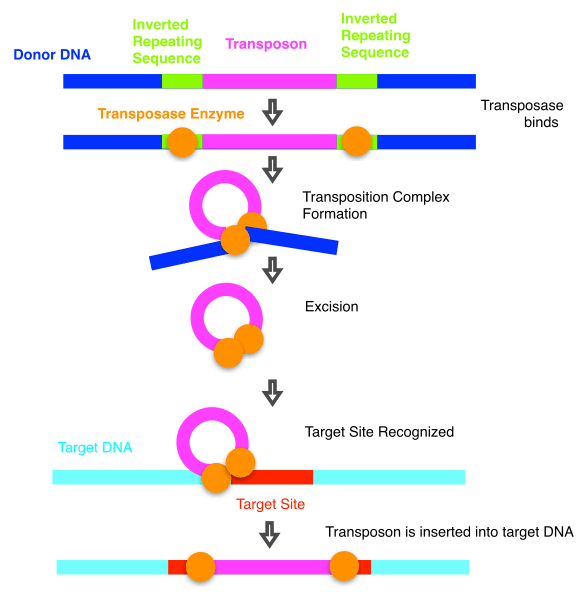
Jumping genes, or transposable elements, are DNA segments that can be cut/copied and pasted from one place in an organism’s genome to another place or places. They’re found in most organisms and comprise a large fraction of many organism’s genomes. Usually, such moves have a negative or neutral effect with the very occasional adaptive mutation. Consequently, large populations tend to filter out active transposons over time. However, eusocial species have a much smaller population size which may be why these transposable elements have been able to accumulate. However, in a ‘chicken and egg’ twist, it’s also possible that having a lot of humping genes also provided the opportunity for adaptive mutations that enabled the transitions to eusociality.
The research continues. Dr. Rubenstein, head of the Columbian lab studying these genomes, notes that such work is answering larger questions at the intersection of behavioral and molecular biology “Understanding how living in complex societies can provide feedback on genome architecture represents an intriguing new area of study that has implications for all types of social animals, perhaps even for humans.”
Ready to read more about shrimp evolution and genomics? Check out these three excellent research articles.
Queller, D., & Strassmann, J. (1998). Kin Selection and Social Insects. BioScience, 48(3), 165-175.
Title Image: Muséum national d’histoire naturelle, CC BY 4.0 https://creativecommons.org/licenses/by/4.0, via Wikimedia Commons

