If you have not already, please read part 1 of our series on genetic variation.
Last time I discussed how genes can be created de novo through mutations in non-coding regions of the genome. De novo genes have been shown to represent a small fraction of the gene pool, although this varies significantly across species. In Drosophila melanogaster, the commonly used fruit fly model organism, researchers recently uncovered 142 de novo genes. This number is substantially more than the ~40 such genes found thus far in humans.

Importantly, it seems likely that many of the de novo genes are functional within an organism. One example of this can be found in the human NCYM gene, a de novo gene that was born during an event that created a novel start site downstream of the MYCN oncogene. The NCYM gene is transcribed in the opposite direction from the MYCN gene (hence the clever name), and is expressed in human neuroblastoma, a type of neural tumor found frequently in children.
Researchers, led by Dr. Akira Nakagawara from the Chiba Cancer Center Research Institute in Chiba, Japan, examined the function of NCYM. Experiments in transgenic mice uncovered a role for the NCYM protein in promoting metastasis from neuroblastoma cells. Thus, the de novo gene NCYM, which only arose relatively recently in humans and chimpanzees, has an important function in regulating disease.
It is likely that scientists will continue to uncover additional de novo genes with important functions, thanks in large part to advances in genome sequencing and computer predictions.
SNPs reveal differences within species
These findings reveal the potential for de novo genes to arise within an organism, and for these genes to develop a function through a combination of sheer probability and selective pressure. What about all of the other changes that occur throughout our genomes but do not produce a definitive protein product? It is estimated that a single nucleotide polymorphism (SNP), a change of an individual DNA building block into a different nucleotide, can occur every 300 bp on average. This would translate into approximately 10 million SNPs within the human genome. Obviously, the majority of these SNPs are benign, but it is also possible for SNPs to play a more functional role in disease.

In addition, SNPs are useful for determining an organism’s haplotype – the combination of inherited alleles on a chromosome. By examining a combination of SNPs across multiple individuals we can uncover links to genetic diseases and map genetic similarities.
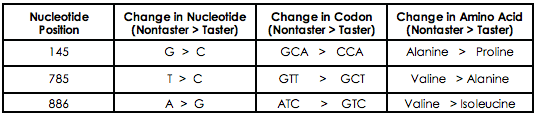
One of the best examples of how a SNP can be examined is through analysis of the TAS2R38 gene. Individuals vary greatly in their ability of taste the bitter compound Phenylthiocarbamide (PTC), which is regulated though 3 different SNPs within TAS2R38. The presence of these SNPS alters the structure of the final protein, inhibiting protein function. It is possible to sequence an individual’s DNA for the presence of these SNPs, although this option can prove relatively expensive. Instead, it is possible to examine the presence of SNPs through restriction enzyme digestion. Restriction enzymes specifically cleave DNA at defined sequences; by taking advantage of the enzyme HaeIII, we can specifically cleave “Taster” TAS2R38 DNA, while individuals with the “Non-taster” SNP will not be digested. Thus, it is possible to quickly screen individuals for the presence of a SNP, which can be easily verified by a PTC taste-test!
To find out more about PTC and the TAS2R38 PCR and restriction digest experiment check out the excellent kits and resources at Edvotek.com.
That does it for part two of my series on de novo genes and genetic alterations. Check back soon as I wrap everything up in part 3!

1 comment
Comments are closed.