In order to communicate their findings to others in their field, scientists will publish their data in peer-reviewed journals. Some of the techniques within these papers have revolutionized the way we do science. Nature, a leading science journal, tabulated the top 100 most cited scientific research articles ever. Many of the techniques within these papers are used in the research lab every day! Here are five of the most common – and most cited – biotechnology techniques that your students can try in your classroom laboratory.
#4 – “DNA sequencing with chain-terminating inhibitors” by F. Sanger, S. Nicklen & A. R. Couslon (1977).
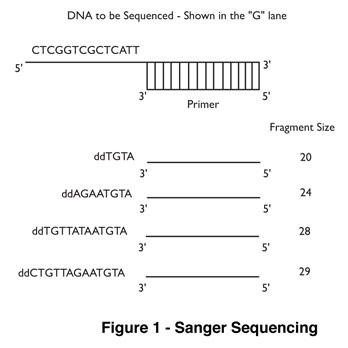
This paper described a technique that determines the nucleotide sequence of DNA by creating a series of copies of the original in vitro. The DNA is mixed with a DNA primer to target the site to be sequenced, the enzyme DNA polymerase I (DNA Pol I), and free nucleotides. DNA Pol I uses the primer to start synthesis of the new strand of DNA in the 5’-3’ direction using the existing DNA as a template. To this mixture, Sanger added dideoxynucleotides (ddNTPs). These nucleotide analogs lack the 3’ hydroxyl group, making it impossible for the polymerase to add another nucleotide to the end of growing strand. This creates a series of DNA fragments of differing size that can be used to map the location of each nucleotide in a given piece of DNA (Figure 1). This technique won the 1980 Nobel Prize in Chemistry and is still being used today. Explore DNA sequencing in your classroom!
Southern blotting is used to identify specific DNA sequences within the context of an organism’s genome. First, the genomic DNA is cut into smaller fragments using restriction enzymes and separated into discrete bands by agarose gel electrophoresis. Next, the DNA is transferred to a nylon membrane. The membrane is incubated with sequence-specific DNA probes that bind to the region of homology in the membrane-bound DNA. Each probe is labeled so that we can easily detect the pattern of hybridization (Figure 2).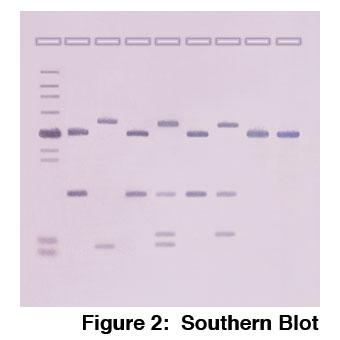
This technique can identify sequence differences, or “polymorphisms”, that are found throughout the genome. For example, differences in the nucleotide sequence affect the number and location of restriction enzyme sites, creating Restriction Fragment Length Polymorphisms (or RFLPs). If the DNA is not digested by the restriction enzyme, the probes will only hybridize to a single DNA segment. If a restriction site occurs within this sequence, the probe will hybridize with multiple bands of DNA.
Because DNA samples from different individuals produce different patterns of bands, scientists can use RFLPs to distinguish between individuals. In fact, early DNA fingerprinting analysis involved RFLP analysis of crime scene DNA. RFLP-specific DNA probes were used to probe a blot that contains crime scene and suspect DNA. A match between a suspect’s DNA and the crime scene DNA suggests that the person was at the crime scene. Try this technique in your classroom laboratory!
Protein molecules pose a unique challenge for electrophoresis because their complex shapes and different charges affect how they migrate through a gel. This problem was solved by SDS polyacrylamide gel electrophoresis (SDS-PAGE). In this technique, the protein’s secondary structure is unfolded using detergent and reducing agent before electrophoresis. This allows proteins to be separated according to their molecular weight, and not by shape or charge (Figure 3). Visit our website to learn about performing SDS-PAGE and other protein experiments. 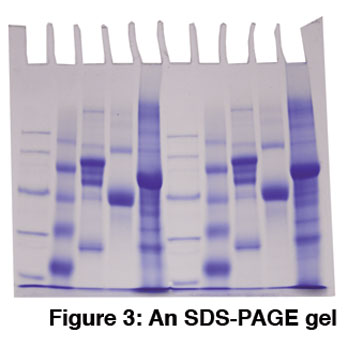
PCR is a technique that allows researchers to quickly create many copies of a specific region of DNA in vitro. Because of its ease of use, this technique has become indispensable in medical and life sciences labs to identify genetic mutations and infectious agents. Since PCR requires very little starting material, it is ideal for forensic analysis of biological samples. For the development of this technique, Mullis won the 1993 Nobel Prize in Chemistry.
 The bestselling EdvoCycler™ has been reimagined to offer the classroom advanced PCR functionality at the lowest sample price. At 48 wells, the EdvoCycler™ 2 doubles the capacity of the original machine and offers superior performance and ease-of-use in a sleek new form factor with a vivid touchscreen display. Get it now at the introductory price of $1499 (subject to change). Be sure to explore PCR experiments to go with your new thermal cycler.
The bestselling EdvoCycler™ has been reimagined to offer the classroom advanced PCR functionality at the lowest sample price. At 48 wells, the EdvoCycler™ 2 doubles the capacity of the original machine and offers superior performance and ease-of-use in a sleek new form factor with a vivid touchscreen display. Get it now at the introductory price of $1499 (subject to change). Be sure to explore PCR experiments to go with your new thermal cycler.
Plasmids are circular pieces of DNA that are used by molecular biologists to study, analyze, and clone genes. In order to be used for research and other scientific purposes, the plasmid has to be isolated from its bacterial host. The alkaline-lysis plasmid extraction technique is a simple and reliable method to isolate the DNA from the bacterial cells.
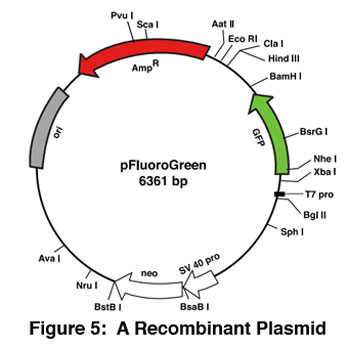
With Edvotek™, you can perform this exact experiment in your classroom!
Original article:
Van Noorden, R., B. Maher & R. Nuzzo. (2014). The Top 100 Papers. Nature 514, 550–553
