DNA is one resilient molecule! In addition to having mastered the arts of copying and repairing itself, its structure is very stable. Under ideal conditions, this molecule can survive for close to a million years, and even when exposed to harsh elements, can still last for several days.
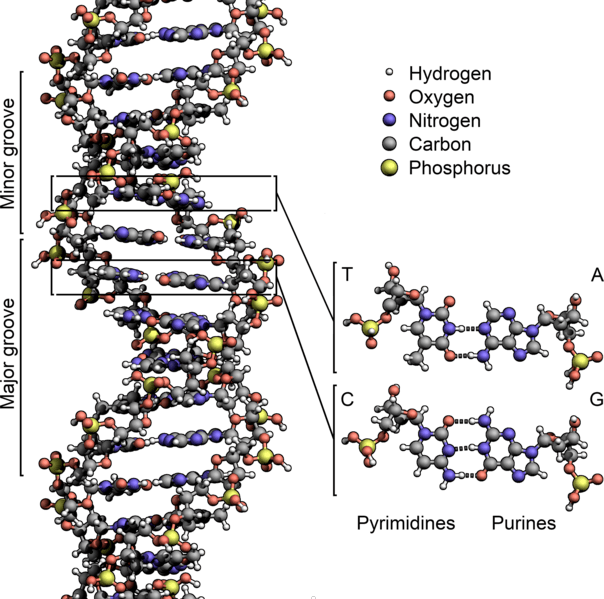
One secret to its toughness is DNA’s double-stranded helix shape. This shape places the molecules most vital parts – the genetically informative nucleotide bases – inside where they are protected from chemical mutagens. Meanwhile, the outside sugar-phosphate backbone is made out of ribose sugars that have lost one oxygen and one hydrogen atom. This small atomic change makes DNA far less susceptible to chemical breakdown due to a reaction with water (i.e. hydrolysis).
This is not to say that DNA is impervious. (So please continue to keep your DNA samples stored in the fridge and/or freezer whenever you can.) DNA does degrade. This is particularly true when the molecule is exposed to sunlight, water, heat, large temperature change, and oxygen.
When DNA degrades it generally changes shape in one of three ways. The first, cross-linking, occurs when two nearby nucleotides form a new covalent bond. The second, deamination, is when one of the four nucleotides (cytosine, thymine, guanine, or adenine) loses an amino group. The third, fragmentation, is when the DNA strand is broken into smaller pieces.
These changes make working with degraded DNA, often called ancient DNA or even just aDNA, challenging but not impossible.
Scientists in the field of paleogenetics or archaeogenetics capitalize on DNA’s durability and on rapidly evolving biotechnologies, in order to study aDNA that has been extracted from all sorts of unexpected places – skeletons, mummies, eighteenth-century plant collections, lake sediment samples, excavated pottery fragments, and ice cores, to name a few!
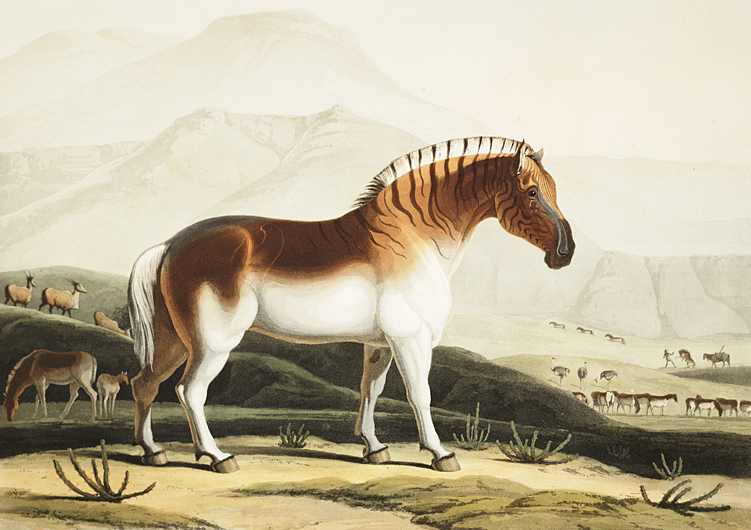
The very first aDNA that was read by scientists came from a quagga – an extinct zebra-like species. In 1984 biochemists from the University of California Berkeley extracted DNA from a 150 years old museum specimen of this creature and successfully sequenced 229 base pairs.
Fast forward 36 years and the first aDNA genome was reconstructed by scientists who extracted DNA from the frozen remains of a man who was estimated to have lived around 4,000 years ago in Greenland. They were able to retrieve around 80% of the genome and identify over 300,000 informative single nucleotide polymorphisms.
Today, the field of paleogenetics is exploding and helping archeologists, paleontologists, ecologists, and evolutionary biologists deepen our understanding of the past and sometimes even rewrite history.
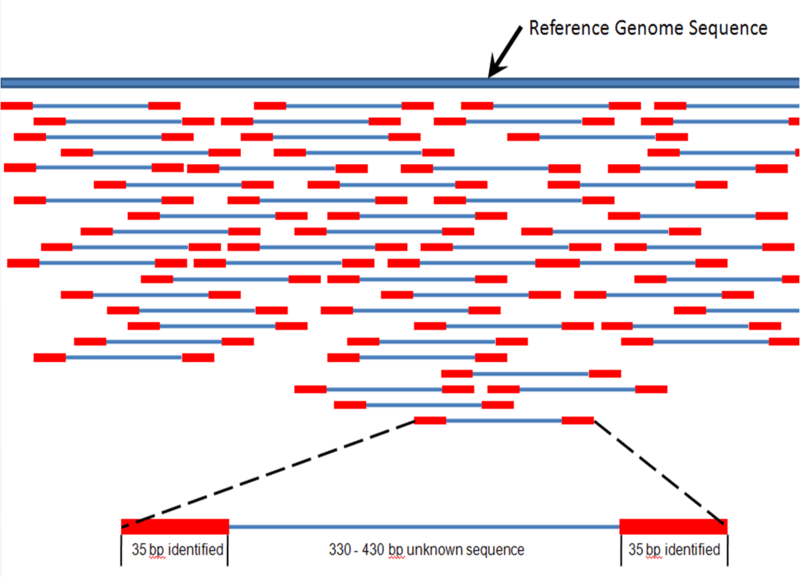
This is due in part to continued advancements in next-generation sequencing technology. This technique rapidly reads large DNA samples by breaking long DNA sequences into smaller fragments, sequencing these fragments, and then using sophisticated computer algorithms to overlay each sequence and reconstruct a genome. Such a procedure naturally lends itself to aDNA which is often fragmented due to exposure and degradation.
Fast computers and sophisticated algorithms are also helping guard against another major stumbling block of aDNA analysis – contamination with modern DNA. Such contamination comes from both humans conducting the research and from microbes in the environment. Traditionally, contamination has been combated by extracting and sequencing aDNA under extreme sterilization conditions. Now, new bioinformatics analyses help scientists to also distinguish between the ancient DNA sample and modern DNA contaminants based on the presence of certain mutations unique to the former.
Another important breakthrough – at least in ancient human studies – is an anatomical discovery. In 2014 scientists found that DNA is stored in incredibly high concentrations (up to 100 x) in a small part of the human skull known as the petrous. Using this much richer source of aDNA scientists have been able to extract DNA from older and more diverse skulls. Notably, they’ve been able to read DNA from ancient human remains in the tropics and subtropics – two key regions in our species history and evolution.
Here are a few of my favorite aDNA studies from the last few years brought to you by a combination of human ingenuity and one very important and sturdy molecule.
- World’s oldest chocolate was made 5300 years ago—in a South American rainforest (2018)
- ‘Viking’ was a job description, not a matter of heredity, massive ancient DNA study shows (2020)
- Social life of extinct sabre-toothed cat revealed by ancient DNA (2020)
- US Government to Repatriate Kennewick Man (2016)
- The world’s first farmers were surprisingly diverse (2016)
- Ancient DNA Reveals Complex Story of Human Migration Between Siberia and North America (2019)
Interested in combining history and molecular biology in your classroom but don’t have a state of the art cleanroom to analyze aDNA or even ethically obtained aDNA samples? Not a problem. DNA extracted from living specimens can also be used to learn about the past. Discover how scientists used ‘modern’ mtDNA samples to determine our species ancient migration routes and then have your students extract and amplify a highly conserved section of their own mitochondrial genome.




2 comments
Comments are closed.