A cell’s DNA contains a wealth of valuable information. This information allows organisms to create the proteins they need to live. Scientists also use the information stored within DNA to answer questions, diagnose diseases, solve crimes, and even unravel Earth’s history. For example:
- Forensic scientists can match highly conserved regions of DNA taken from crime scene samples with a suspect’s and submit the results as evidence in criminal cases.
- Doctors analyze DNA to identify genetic disorders like Cystic fibrosis and Huntington’s disease or to flag genetic predispositions to certain diseases like cancer and heart disease.
- Virologists and microbiologists use sequences to detect certain organisms in a sample or to broadly describe the microbial community present.
- Evolutionary biologists, ecologists, and phylogeographers study patterns of DNA nucleotide changes to reconstruct the past and determine how life has evolved on Earth and also how different species have responded to changing environments.
Likely, you are already familiar with several of the above examples. You may have read or heard about them in the news, studied them in a textbook, or performed an experiment relating to one or more of them. However, ecological applications of DNA technologies are less well known and only just beginning to be taught alongside the more traditional forensic and medical applications. A great example of the growing intersect between genetics and ecology is the field of phylogeography.
Phylogeography 101
Phylogeography is the study of historical processes that create the past and present geographic distributions of different genealogical lineages. This field has grown rapidly in the past two decades thanks in part to newly developed sequencing technologies. Today phylogeographic studies are used to help us better understand all types of species from bacteria and viruses to elephants and whales. Popular genetic testing kits that help individuals discover their ancestral roots are also rooted in phylogeography. (As of 2020 approximately 30 million people and counting have used these services.) Finally, phylogeography is helping scientists study the ecological effects of past climate changes and helping conservationists protect and preserve threatened species.

In phylogeography, information about differences in the DNA of individuals (i.e. a species’ genetic variation), the geographic distribution of this variation, and the evolutionary relationships between different DNA sequences are analyzed together to detect genetic signals of past events. Scientists begin by collecting DNA samples from many individuals throughout the species’ current range – and sometimes from fossils or museum samples. Each sampled individual is then assigned to a DNA group, or haplotype, based on his or her genetic profile. This haplotype is linked back to a location based on where the sample was collected or where the individual was born. Researchers then group these samples into geographic populations.
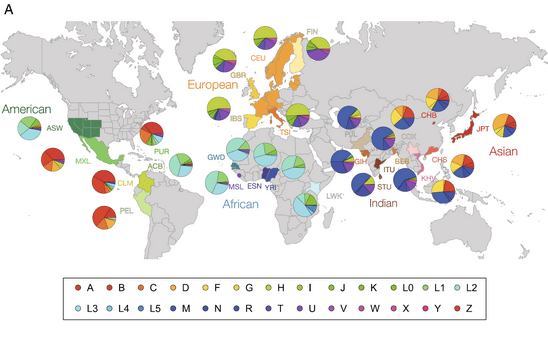
Phylogeographers pay particular attention to which haplotypes are unique to a single population and which are shared by many populations as well as the number of haplotypes observed in each population. This information is described using mathematically based measures such as effective population size, expected heterozygosity, and F-statics. It is also visually summarized by pie graphs of each population’s haplotypes which are added to a map at each population’s location. Together, population genetic statistics and haplotype maps describe the genetic patterns of a species or the “geography” part of its phylogeography.
To predict what events created the observed genetic patterns, scientists next reconstruct the evolutionary history of all the different haplotypes using phylogenies – the “phylo” part of phylogeography. Phylogenies describe patterns of ancestry. Often phylogenies are displayed as trees or networks where whatever is being compared – family members, species, or in this case unique haplotypes – are on the tips of each branch. Common ancestors are represented by the intersection between two branches which are known as a node. Haplotypes that are several nodes away from one another are more distantly related than those that are only separated by a single node. Similarly, haplotypes that are connected by short branches are more closely related than those connected by longer branches. The tip of each branch is labeled with the haplotype’s name and in some cases an additional pie chart showing all the populations where the haplotype can be found.
By combining data from phylogenies and population genetics scientists can reveal key events in the history of a species. Phylogeographical studies can help scientists determine when/where members colonized new areas (expansion events), when/ where populations went through a sudden decline in size (bottleneck events), when/where populations divided and became isolated from each other (vicariance events), and when/where members from two different populations met (admixture events). Once a species’ phylogeography has been created, the genetic information can then also be organized and shared in a database called a GRDB (Genotype Reference Database) that can be used to predict the geographic origins of an individual.
Teaching Resources
Discovering an individual’s or a species’ history using phylogenetic analysis can be a great way to explore DNA, mutations, population genetics, and phylogenies and master biotechnologies such as sequencing, electrophoresis, and restriction enzyme or SNP analysis. Here are three resources to bring this subject into your classroom.
- The DNA testing company 23&Me has several lesson plans for students from elementary school through undergraduate, that range from a guide to talking about family history with relatives to analyzing SNP data sets.
- The Tree of Life website has a Phylogeography section with classroom activities, a quiz, and useful links.
- Our experiment Safari Family Reunion immerses students in the technologies and applications of phylogeography by performing electrophoresis on the DNA samples of two lion cubs, analyzing phylogenic trees and haplotype maps, and recommends a conservation plan for the cubs.


1 comment
Comments are closed.