Synthetic biology, the science of redesigning an organism to accomplish a new activity or to possess a beneficial trait, has become an indispensable part of modern biotechnology. We’ve written before about the amazing synthetic biology projects that are already changing our lives, but researchers are continuing to push the boundaries as new techniques become available. For example, scientists are using synthetic biology to combat climate change, produce revolutionary new materials, and even to create novel organisms. This week, I wanted to write about some incredible new research that is modifying bacteria one nucleotide codon at a time.
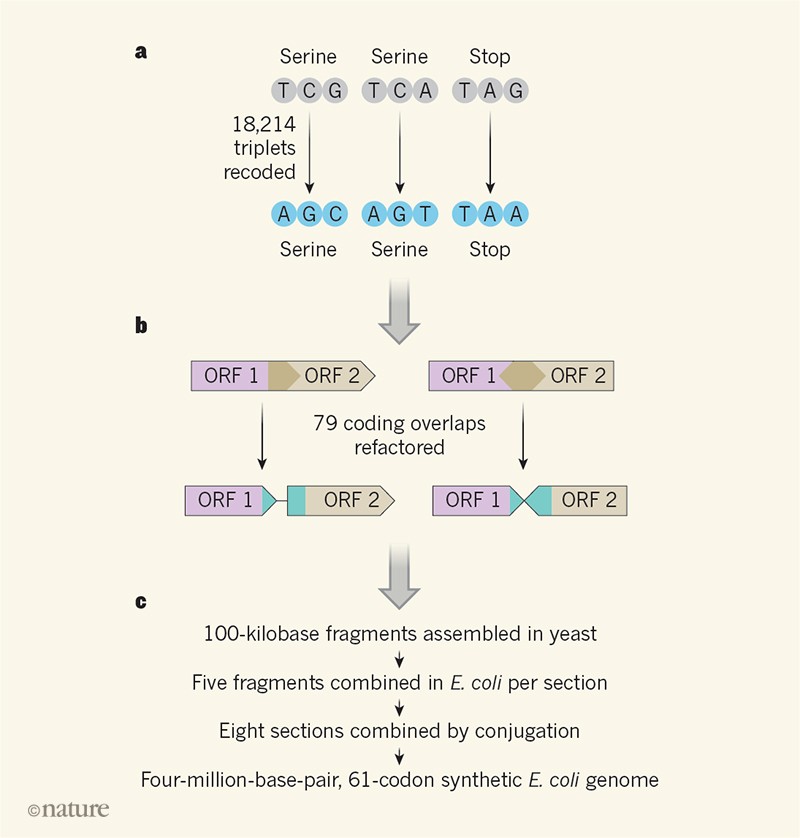
A few years ago, a group of researchers created a synthetic E. coli genome that was able to use fewer codons than normally found in nature. Wild-type E. coli use 64 different codons – combinations of 3 nucleotide bases – to encode 20 amino acids. This system means that each amino acid can be encoded by multiple different codons. For example, the amino acid Serine is encoded by six different codons (TCG, TCA, TCT, TCC, AGT, and AGC). In the synthetic bacteria, the researchers (led by Dr. Julius Fredens in the lab of Dr. Jason W. Chin) were able to remove 3 of these codons from the entire genome. This included two codons that encode Serine, TCG and TCA, and the TAG stop codon. The researchers carefully identified every instance of these codons in the E. coli genome and then systematically replaced them with a new codon for the same amino acids. In total they replaced 18,214 codons throughout the genome! Amazingly, the researchers were able to demonstrate that the bacteria could live while using only 61 codons.
Earlier this week Dr. Chin’s lab, this time lead by Wesley E. Roberson, took their synthetic E. coli bacteria a step further by removing the tRNA molecules that recognize the TCG and TCA codons. These new bacteria can no longer recognize any segment of DNA that contains those codons. Fortunately for the bacteria, since they no longer contain either codon they are able to grow uninhibited. However, the viruses that normally infect the E. coli were not so lucky. It turns out that the viral genomes still contain many of the TCG and TCA codons; since the bacteria can no longer recognize these codons the viruses can no longer replicate. In short, the modified bacteria are now resistant to viral infection!
Finally, the researchers were able to program the bacteria to use artificial tRNAs. Since they knew that the TCG and TAG codons were no longer being used by the bacteria, the researchers instead used those codons to produce chains of synthetic polymers that are not found in nature. Future studies will hopefully determine if the bacteria can even be used to produce antibiotics and other drugs! As an added bonus, the viral resistance makes the E. coli a perfect strain for industry use where a viral infection can ruin thousands of liters of bacterial cultures.
These studies from Dr. Chin’s research laboratory reveal some of the potential for synthetic biologists to not just modify aspects of an organism, but instead completely rewrite the genetic code. If you’d like to explore synthetic biology in your classroom, Edvo-kit #331 – Investigating Synthetic Biology – offers an exciting look at the ways that bacteria can be turned into living factories. Students will engineer a plasmid and insert it into E. coli, prompting the bacteria to produce an enzyme responsible for the “wintergreen” odor!
Note: the articles cited in this post require a subscription to access and can be found here:
- Total synthesis of Escherichia coli with a recoded genome:
- Sense codon reassignment enables viral resistance and encoded polymer synthesis:
Featured image: “Colorized scanning electron micrograph of Escherichia coli, grown in culture and adhered to a cover slip”. National Institute of Allergy and Infectious Diseases (NIAID).

