Taq polymerase is a powerhouse molecule of biotechnology! Daily it’s used to solve crimes, diagnose diseases, personalize medical treatments, monitor food, and power synthetic biology creations – to name just a few. In 1989 it was named Molecule of the Year by Science Magazine and its prevalence has only increased since then. Today Taq is found on the shelves and freezers of millions of labs and classrooms waiting to help us examine and amplify DNA.
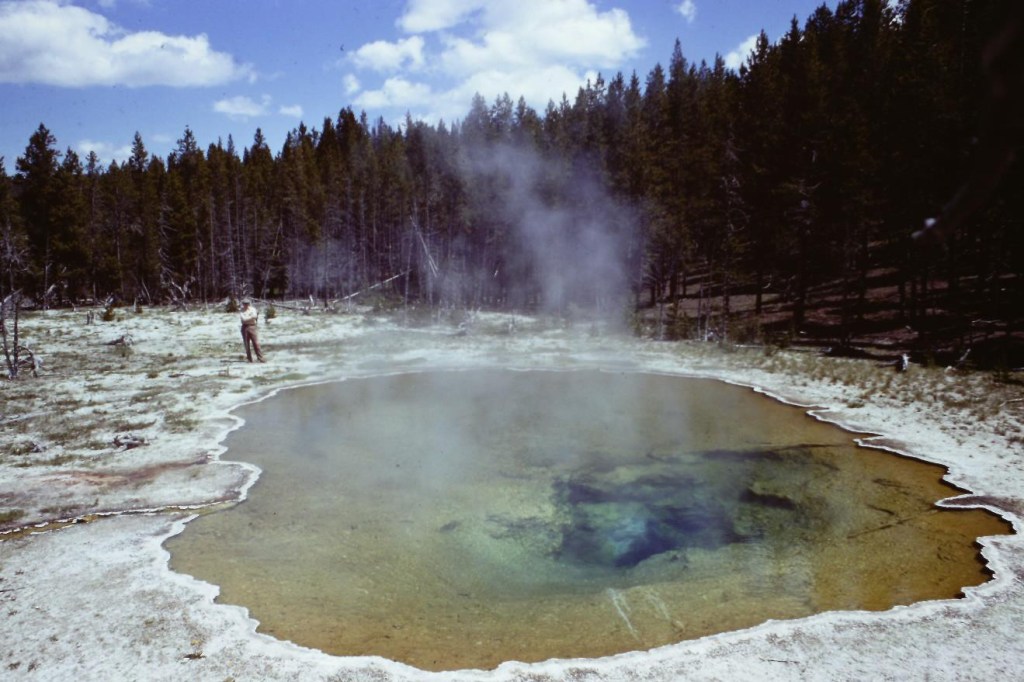
However, the story of Taq does not begin in a lab but in the wilderness of Yellowstone National Park. There Dr. Thomas Brock was studying the microbial ecology of natural hot springs – trying to figure out how certain bacteria can survive at such high temperatures. In 1966 at Mushroom Springs (pictured right) he collected a pink filamentous microbe that was thriving at a sweltering 73oC (163.4oF).
Dr. Brock bought the discovery back to his lab and after months of trial and error, was eventually able to culture the heat-loving microorganism. This was thanks in part to the hard work of an undergraduate named Hudson Freeze. Brock and Freeze named the bacteria Thermus aquaticus and reported the discovery in the Journal of Microbiology in 1969. They also sent a sample of T. aquaticus to the American Type Culture Collection where it would be freely available to anyone interested in studying the thermophile.
And there were several interested groups. In the 1970s biochemists from both universities and private industries were on the hunt for proteins that would not be destroyed or deactivated at high temperatures. Where better to look than heat-loving microorganisms? In 1976 (ten years after the bacteria was first discovered) a master’s student Alice Chein along with her advisers successfully extracted several proteins from samples of T. aquaticus that they obtained from the public cell culture collection. Among them was a small polymerase (~94 kDa) which they named Taq polymerase.
Polymerases are enzymes that catalyze synthetic reactions – that is they cause or accelerate the assembly of large polymer chains from monomer units. In the case of Taq polymerase, the units are nucleotides and the final polymer DNA. Whenever a cell divides DNA polymerase is needed to allow the cell to replicate its DNA. The cell does this by unzipping the DNA into two single strands and then using polymerases to help add nucleotides to the 3 prime end of a template strand.
Every living organism in the world, and even some viruses, have at least one DNA polymerase enzyme in their repertoire and many have several. This means that DNA polymerases are a large and essential family of enzymes! However, the newly identified Taq polymerase had some unique properties. For one it was easily isolated from other enzymes. This meant that it could be produced in a purer and more potent form than most other polymerases. It also had high efficiency and amplification capacity. The former refers to how effectively the enzyme grabs onto and uses nucleotides – this minimizes how many extra nucleotides are needed for the reaction. The latter is the rate at which the polymerase strings together nucleotides. Taq can add ~100 nucleotides per second! However, the ‘pièce de résistance’ was that Taq polymerase was stable at high temperatures. It would take another decade for scientists to match these particular strengths to a novel biotechnology called PCR. However, the resulting pairing revolutionized the life sciences.
In 1985 Dr. Kary Mullis invented a way to create billions of copies of a specific region of DNA in vitro. The procedure, called the Polymerase Chain Reaction (PCR), involves combining a small amount of the DNA of interest with primers, nucleotides, and a polymerase and then exposing the mixture to a cycle of three different temperatures. However, this technique was slow and labor-intensive because scientists needed to add fresh enzyme after each temperature cycle. This was because the initial enzyme used, E. coli DNA polymerase I, was degraded by the high temperatures and large temperature changes of each cycle. In 1988 Mullis and his colleagues at the Cetus Corporation revised their PCR protocol (and rebuilt the temperature-changing thermocycler that the procedure depended on) to work with Taq polymerase. The revision was an instant success. Not only could PCR now be automated but the new cycling times made the whole process much faster. In addition, the higher temperatures of Taq PCR also increased the primer specificity which significantly reduced the production of nonspecific products.
Over thirty years later Taq continues to dominate the PCR world although a few thermostable alternatives now exist such as the polymerases found in Pyrococcus furiosus (Pfu) and Pyrococcus woesei (Pwo). One weakness of Taq is that, unlike most polymerases, Taq can only help build in the 5’ to 3’ direction and cannot proofread in the 3’ to 5’. This means that it cannot detect and correct mismatched nucleotides which leads to a higher error rate (somewhere between 1.1 x 10-4 errors and 8.9 x 10-5 errors per base pair per duplication.) Another drawback is that Taq is still active (although slow) at lower temperatures which can lead to primer dimers. In both these cases, combining Taq with another enzyme has proved an effective solution. High-fidelity polymerases combine Taq with additional proofreading polymerases while Hot-Start Taq mixes Taq with inhibitors that allow the former to work only at higher temperatures.
Taq is an incredibly powerful molecule that has opened the flood gates to a dizzying array of discoveries and life-changing technologies. The discovery and development of Taq also illustrates the power of diverse scientific research. Taq is available to researchers today thanks to the hard work of scientists asking a range of ecological, biochemical, and biotechnological questions and thanks to major contributions from everyone from undergraduate researchers (Hudson Freeze) to big-business science think tanks (Cetus Corporation).
Dive deeper! Learn more about Taq polymerase and PCR at our learning center or try one of our classroom PCR experiments.
Title Image Credit: Jawahar Swaminathan and MSD staff at the European Bioinformatics Institute, Public domain.



