SNP analysis of the PTC gene using the Polymerase Chain Reaction (PCR) is one of my favorite ways to demonstrate the link between genotype and phenotype in the classroom laboratory. This wonderful experiment involves the isolation of human DNA from the students, amplification of the TAS2R38 gene by PCR, restriction enzyme digestion, and agarose gel electrophoresis. Finally, students are given the opportunity to directly compare their gel electrophoresis results to the ability to taste PTC, solidifying the link between genotype and phenotype.
Unfortunately, the pandemic has made it significantly more difficult to perform many in-person experiments, including SNP analysis of the PTC gene using PCR. Here are some of the most frequently asked questions from the past year, as well as some general tips and tricks to ensure you get the best results possible.
Q. How can we perform the DNA extraction without requiring the students to isolate cheek cells?
A. DNA isolation from human cheek cells has been the preferred method in the past, but due to the pandemic we advise having the students collect DNA from hair follicles. You can find a simple protocol here. Although the hair extraction takes slightly longer than with cheek cells, the results are nearly identical and it can be performed safely while masked.
Q. The hair DNA extraction worked great, but we don’t have time to run the PCR the same day. What should we do with the student samples?
A. The experiment is designed to easily pause between any of the modules. After isolating the student DNA it can be frozen for up to two weeks before performing the PCR. Similarly, after the PCR amplification or restriction digest protocols have been completed the samples can be refrigerated overnight or frozen for up to two weeks. Finally, if you would like to save the agarose gels we have an excellent guide that you can find here.
Q. My classes have been split into multiple sections. Is there a good way to divide the reagents between periods or days?
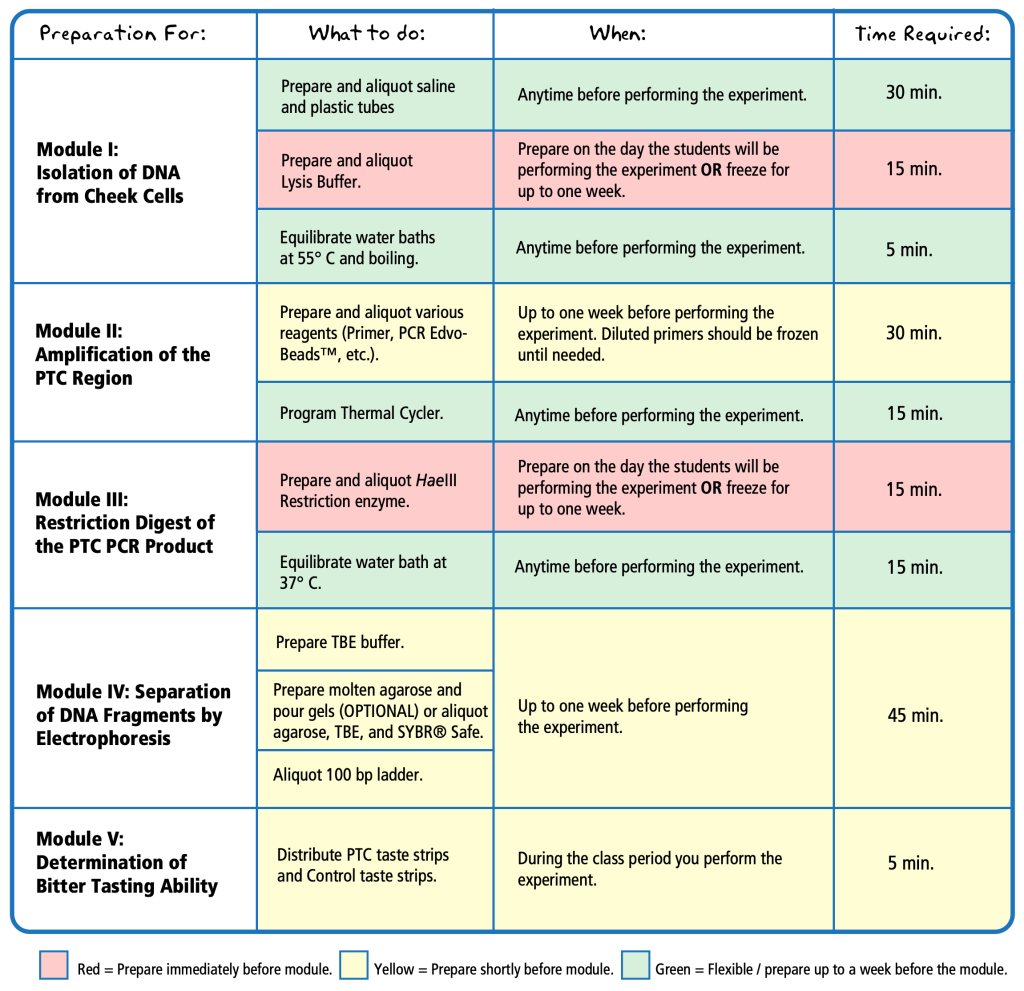
A. The experiment is very easy to modify for split periods or emergency closures, simply follow the instructions on the first page of the instructor’s guide. For the DNA extraction, prepare the lysis buffer as instructed, aliquot into tubes for the students, and then immediately freeze any that will not be used immediately. The PCR reagents in Module II can be stored for up to a week in the refrigerator, or frozen for longer. Finally, the Restriction Enzyme digest reagents should be prepared, aliquoted, and frozen as with the DNA extraction materials from earlier.
We have many additional tips for this experiment on our website at https://www.edvotek.com/learning-center. Be sure to check out our instructional videos, including DNA isolation and PCR. We also have an excellent troubleshooting guide to DNA extraction and PCR here. Finally, if you have any other questions don’t hesitate to contact us!


1 comment
Comments are closed.