Deoxyribonucleic acid, or DNA is the genetic material found in living organisms from bacteria to plants to humans, and even some viruses. DNA is the blueprint used to build an organism. The directions coded for by our DNA controls everything from growth and development to cell specification, neuronal function, and metabolism.
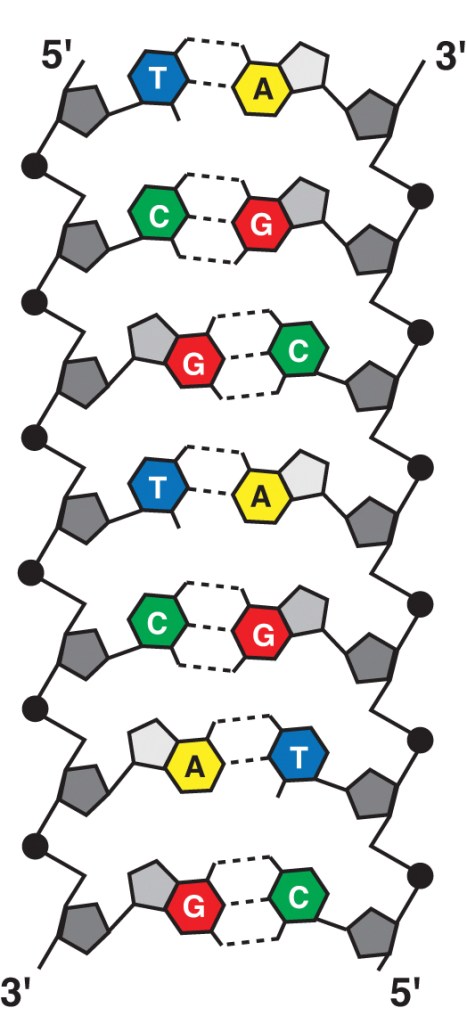
DNA is built from the nucleotides Adenine, Thymine, Cytosine, or Guanine. The nucleotides are linked to one another, forming long strands of DNA. The specific order of the nucleotides creates genes, which are the units of heredity. Many genes direct cells to create proteins, which have many unique functions with the cell. Other genes produce important RNA molecules like transfer RNA and ribosomal RNA.
Each strand of DNA is paired with a second strand following specific rules: A can only pair with T, and C with G. Hydrogen bonds between the paired nucleotides keep the two strands of DNA together as it is packaged into chromosomes.
The DNA in the human genome is wrapped up into 23 pairs of chromosomes. Although most of this DNA is identical between individuals, small sequence differences, or “polymorphisms”, occur at specific locations throughout the genome known as loci. These differences are passed from parent to child, and segregate independently following Mendel’s Laws of Inheritance.
There are many types of DNA sequence differences found within the human genome. The simplest is the single nucleotide polymorphism, or “SNP”. This is a locus where the DNA sequence differs by a single letter. This means that if we zoom in on a single nucleotide within a genome, most people might have an A at that location, but a small percentage of people have a T.
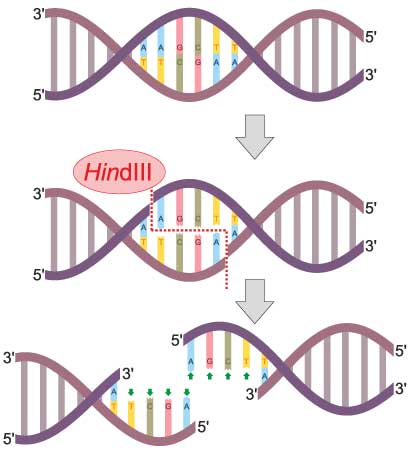
Now, some of these SNPs create Restriction Fragment Length Polymorphisms, or RFLPs. So what does this mean? A restriction endonuclease is a highly specific enzyme that acts like a molecular scissor, cutting DNA into fragments based on its exact sequence. SNPs can create or destroy restriction enzyme sites based on their location within the genome.
For example, the restriction enzyme Hind III cuts DNA with the sequence AAGCTT. Imagine there is a SNP in the restriction site at the second A, where this nucleotide can be an A or a T. Hind III can only cut when the DNA has an A in that location. If we digest the DNA with HindIII, and analyze it using agarose gel electrophoresis, we’d be able to identify the sequence of our sample at that location based on whether the enzyme cut or not.
Another type of RFLP is the Variable Number of Tandem Repeats, or VNTR. A VNTR is a short repetitive DNA sequence. The sequence repeat itself is generally 15-35 nucleotides long, though they can be as short as 2 base pairs and as long as several thousand base pairs. A VNTR can occur between five and 100 times at a specific location.
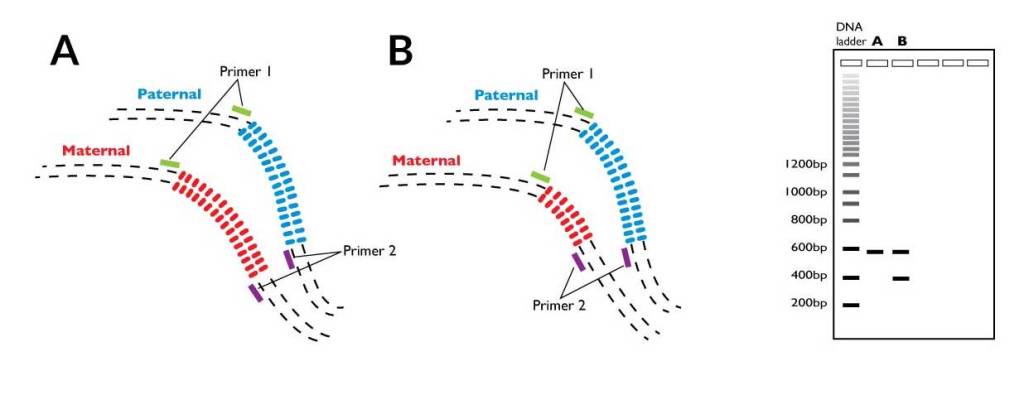
VNTRs change the restriction digest pattern by changing the distance between two enzyme sites. Here, we have VNTRs, represented by colored ovals, between two primer sites. After PCR and gel electrophoresis, the banding patterns will look unique because there are different numbers of nucleotides in that spot.
The location of many SNPs, RFLPs, and VNTRs have been mapped to specific locations within the human genome. These polymorphisms exist at a certain rate within the human population. For example, there is a SNP that occurs in chromosome 19. Nearly 90% of the global population has the G allele, and about 10% of the population has the A allele.
Because each allele is common within a population, we can’t use a single polymorphism to distinguish between two people – the likelihood of them having the same allele is high. However, we can use biotechnology techniques like PCR and DNA sequencing to analyze multiple loci at one time. This allows us to create DNA profiles that describe a specific person.
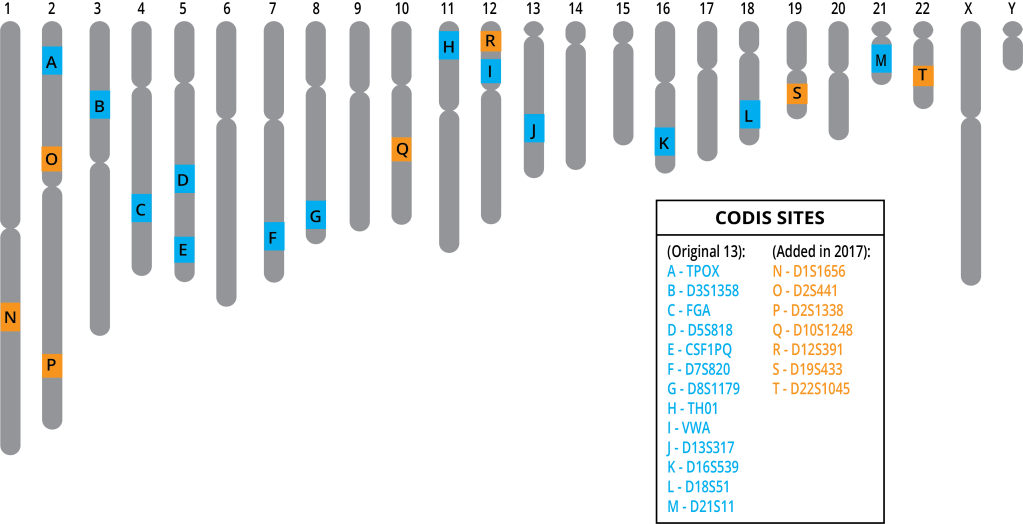
DNA profiles can be used to determine parentage, to discover missing people, and to identify potential suspects. In the United States, forensic scientists create DNA profiles using 20 specific loci within the human genome. These profiles are compared to profiles within the FBI database called the Combined DNA Index System, or CODIS. If there is a “hit”, or a perfect match, the information is provided to law enforcement.
DNA profiles from crime scene evidence provide strong evidence that a suspect was at a crime scene, but that data alone cannot convict a person of a crime. Many lines of evidence, including witness statements and alibis, must come together to build a case against a suspect. All the evidence is used to build a case in the court of law.