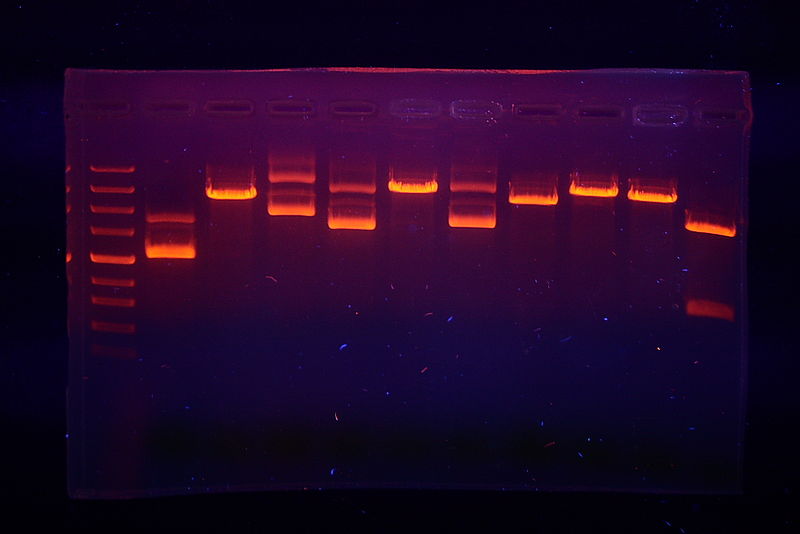
So you’ve selected your plasmid and restriction enzymes, carried out a bunch of digestion reactions, and successfully run and stained a beautiful electrophoresis gel. But now how do you go from this?
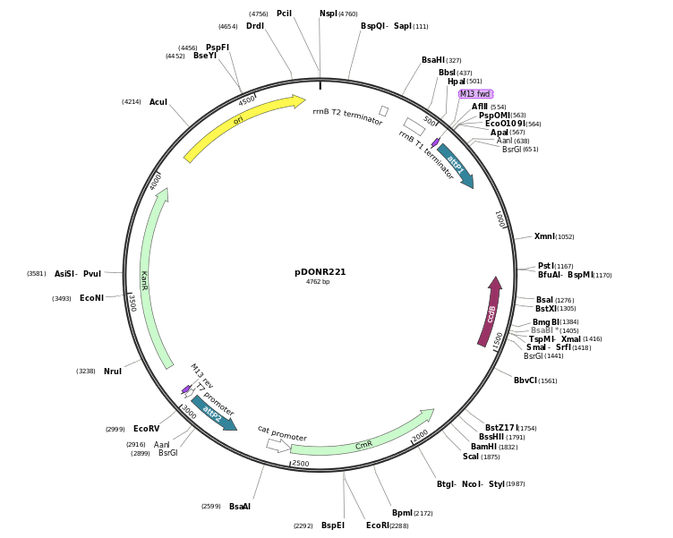
To this?
The answer is by mapping!
Restriction enzyme mapping is a way to describe a plasmid by showing the relative positions of multiple restriction enzyme target sequences. It’s an essential skill in many research and industrial settings. It’s also a great way to get students deeply thinking about DNA and visualizing this molecule and its nucleotide sequences.
Below we’ve outlined some of the key steps, tips, and tricks to make a plasmid map. Keep in mind that mapping is very much a trial-and-error process. Our best tip: treat the data like pieces of a puzzle and don’t be afraid to arrange and rearrange until you find the right fit! Also, remember that there may be more than one ‘correct’ plasmid map for the data.
1) To start, focus on the single enzyme digests. These tell you how many cut sites (also known as cleavage sites) each enzyme has. In plasmids, an enzyme that has one cut site will produce a single fragment. Such one-cut enzymes are extra useful if you’re working with an unknown plasmid as the size of the single fragment is also the size of the plasmid. Enzymes with more cut sites will produce multiple fragments. For example, an enzyme that cuts the plasmid at two sites will produce two fragments.
2) Add the fragment lengths produced by each single enzyme digestion to double check your experiment and make sure that your chosen conditions allowed for complete digestions. Every single enzyme digest should add up to the same final plasmid size. Digestions that sum to a different number are either the result of incomplete digestions or may have two fragments of the exact same size. (Two fragments of the same size will look like a single band on an electrophoresis gel). Summing fragments across all the single enzyme digestions also lets confirm plasmid size even in the absence of a single cut site enzyme.
3) Once you know the size of your plasmid, use a pencil to draw a circle with the total bp number next to it. Take a second to imagine zooming in on this circle’s line to a focus where each point along the circumference is a box with a nucleotide base inside of it and a position number. The ‘box’ at the very top of the circle is given the position number 0. Just to the left of it is the final nucleotide which is at position 0+N where N equals the total size of your plasmid.
4) Begin by mapping the single enzyme digest with the most bands. (Choosing the most bands isn’t required but it tends to make life easier.) Arbitrarily assign the first restriction enzyme cut site of this enzyme to the zero box at the top of the circle. By convention cut sites are represented by a perpendicular line and labeled with the enzyme name. Also write it the the position of the cut site next to the name in parenthesis. Place the next cut size X spaces to the right of the 0/start space where X equals the size of the smallest fragments. Draw the next site X+Y spaces from the 0/start space when Y equals the size of the second smallest fragment. Continue until you’ve accounted for all the fragments. It’s also useful to label the distances between cuts sites (i.e. the numbers X and Y) on the side.
5) Examine the other single enzyme digestions. If you’re new to mapping it can be extremely helpful to create a plasmid map for each individual enzyme digestion and then combine them in a separate step. In many cases, there may be multiple possible locations for the cut sites of different enzymes and no way to know which is the ‘correct’ one until you incorporate information from the double or even triple enzyme digestions. However, having a rough sketch of a possible cut site pattern for each separate enzyme can still make mapping easier.
6) Use combination digestions to determine the placement of one enzyme’s cut sites in relation to the other enzymes’ cut sites*. Combination digestions are the reactions where two or more enzymes were included. Start with a combo that includes the enzyme from step 4. Cross out the fragment lengths in the combination digestion that are already present in the map you drew during this step. Next draw in the additional enzyme’s cut sites in a way that creates the remaining fragments. This is a trial-and-error process and a great place to use a pencil and have an eraser handy!
* Even here there may be several possible ways to draw all the different cut sites. These ambiguities may be resolved as you tackle additional combination digests or may require further experimentation. Keep in mind that the goal is to create a map that accurately reflects the current experimental data which means that one group’s final and ‘correct’ map may be different than another group.
7) Tackle the remaining combination digestions until you have a single plasmid map with all of the enzymes’ cut sites drawn and labeled. This map shows the relative location of all the restriction enzymes used in the experiment. Make sure it’s correct by confirming that all of the DNA fragments created during your digestions are accounted for on the map.
8) You may want to redraw the final map as your map from step 7 will likely have many erased and rearranged lines. In some cases, you may also want to draw a more precise map where the bp position of each cut site is not just an approximation. In this case, divide 360 by the size of the plasmid to determine degrees per bp or kbp and then use this special graph paper to draw in the sites.
Want to see this all in action? Here’s great video.






3 comments
Comments are closed.