 Transformation is a biotechnology technique that is used daily in research laboratories around the world. This technique allows scientists to force E. coli to take up small, circular pieces of DNA called plasmids from the surrounding environment. Using recombinant DNA technology, these plasmids are engineered to contain genes from different sources. Once transformed into bacteria, the plasmids turn the bacteria into living factories to create medications, vitamins and other useful products. For example, scientists use bacteria to make insulin, the medication used to treat diabetes. In the classroom, we can program E. coli with a jellyfish gene, making them glow bright green!
Transformation is a biotechnology technique that is used daily in research laboratories around the world. This technique allows scientists to force E. coli to take up small, circular pieces of DNA called plasmids from the surrounding environment. Using recombinant DNA technology, these plasmids are engineered to contain genes from different sources. Once transformed into bacteria, the plasmids turn the bacteria into living factories to create medications, vitamins and other useful products. For example, scientists use bacteria to make insulin, the medication used to treat diabetes. In the classroom, we can program E. coli with a jellyfish gene, making them glow bright green!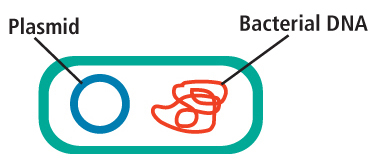
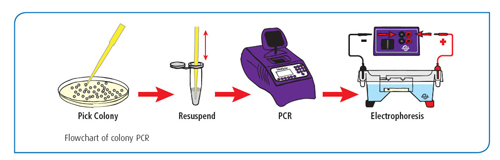
The standard GFP transformation experiment can be enhanced in an extension activity that uses the polymerase chain reaction (or PCR) to confirm the presence of plasmid DNA in the bacterial host. Colony PCR is a high-throughput method used to detect a recombinant insert using the DNA within the transformed cells as a template. In brief, a sterile pipet tip is used to collect a small amount of cells. These cells are resuspended in a PCR mixture containing enzyme, primers, and Taq DNA polymerase. The mixture is heated to 94°C, which lyses the cells and denatures the DNA for the first round of PCR. After the cells are denatured, PCR proceeds as normal (learn more about PCR here). The PCR samples are analyzed by agarose gel electrophoresis. Any cells containing the GFP plasmid should give positive results, proving that the newly acquired genetic information is present in the cells.


1 comment
Comments are closed.