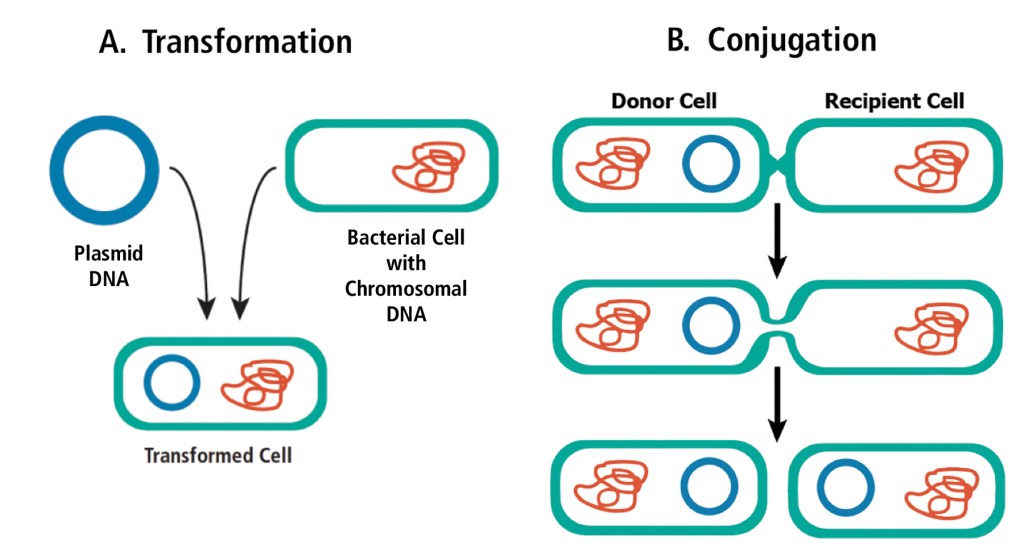
In nature, bacteria pass small pieces of DNA back and forth in two unique ways, which we call transformation and conjugation. In transformation, a bacterium takes up free DNA from the surrounding environment. Some bacteria, like H. influenzae, do this naturally to acquire new DNA. Other bacteria, like E. coli, can be forced to take up DNA and become transformed, even though they are not naturally competent. In contrast, conjugation relies upon direct contact between two bacterial cells. A piece of DNA is copied in one cell (the donor) and then is transferred into the other (recipient) cell through a bridge-like connection. In both cases, the bacteria have acquired new genetic information that is both stable and heritable. Although not essential for the cell’s survival, this DNA can provide an advantage. For example, the gene that codes for beta-lactamase, the enzyme that provides resistance to the antibiotic ampicillin, is found on a plasmid can be passed between bacteria. This gene impacts the way doctors treat people with specific diseases.
In the laboratory, we can engineer plasmids to contain genes from different sources. Once transformed into bacteria, the plasmids turn the bacteria into living factories to create medications, vitamins and other useful products like insulin, the medication used to treat diabetes. In the classroom, we can program E. coli with a jellyfish gene, making them glow bright green!
Since E. coli are not naturally competent, we can heat shock the cells to force them to take up plasmid DNA. In a heat shock transformation, the cells are treated with calcium chloride to make them “competent”. DNA is added to the cells before they are “heat shocked” — or moved quickly between two very different temperatures. It is believed that the combination of calcium chloride and the rapid change in temperature changes the permeability of the cell wall and membrane, allowing DNA molecules to enter the cell. The cells are plated on selective media and allowed to grow overnight. Each colony on the plate represents one transformed cell.
In practice, transformation is highly inefficient—only one in every 10,000 cells successfully incorporates the plasmid DNA. However, since a billion cells are used in a transformation experiment, only a few cells must take up the plasmid to achieve a positive outcome. We can use the data from our experiment to calculate the transformation efficiency, or the number of cells transformed per microgram of plasmid DNA. To calculate the transformation efficiency, we need a few pieces of data:
- The amount of the plasmid that is used in the transformation, in micrograms. A common amount of plasmid to use for transformation is 10 nanograms, or 0.01 micrograms.
- The final volume of the cell suspension at recovery. This includes the volume of the competent cells and the recovery broth.
- The volume of cells plated on the nutrient agar. Since most of the time we will not plate the entire volume of transformed cells, this will allow us to determine the proportion of cells that were plated.
- The number of colonies on the plate. Each colony represents one transformed cell.
The data is then plugged into the below equation to determine the number of transformants per microgram of DNA. Transformation efficiency generally ranges from 1 x 104 to 1 x 108 cells transformed per μg plasmid.
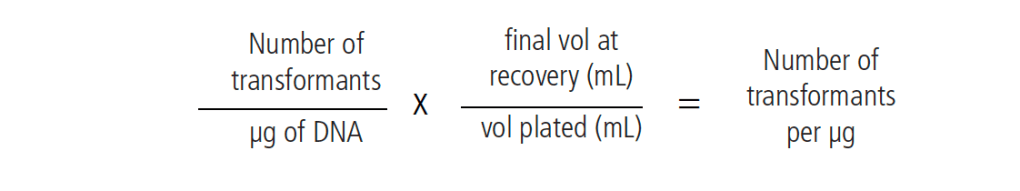
Want to try the transformation efficiency calculation before performing this experiment? Our downloadable worksheet and video make a perfect pre-lab lesson! Try it and let us know what you think!



5 comments
Comments are closed.