This post is for you if:
- You’re looking at a DNA sample tube of clear liquid and wondering “Is there really anything in there?”
- You’re trying to optimize a PCR, sequencing, cloning, or transfection experiment.
- You love the subject of lab book math!
- You involuntarily shudder when you see A260 or A280.
Whichever category you fall into (we won’t judge), read on.
DNA Quantification 101
Following DNA extraction and/or purification researchers will determine the concentration, yield, and, if possible, purity of their product. Such quantifications are a great gauge of the success of these procedures and the integrity of the starting material. They also provide useful information if downstream experiments (like sequencing, PCR, or cloning) go awry and require troubleshooting. There are four major ways to quantify DNA: UV absorbance, fluorescent measurements, agarose gel electrophoresis, and qPCR. By far the most common is UV absorbance.
The Lab Nitty-Gritty of Measuring DNA with UV Absorbance
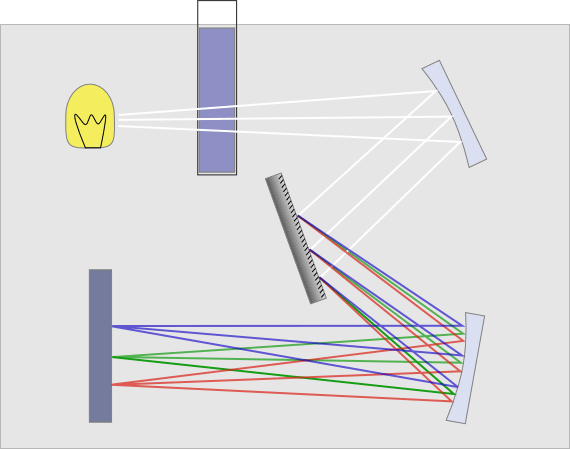
DNA quantification by absorbance requires a UV spectrometer. This is a machine that measures the intensity of light transmitted through a sample. Some UV spectrometers are made just for DNA and RNA quantification. They’re often called drop spectrometers because they usually require a ‘drop’ of the sample between 1 mL and 5 mL. General lab UV specs also work well at quantification. They just require diluting a subsample of the DNA to a volume that can fill the cuvette. Both machines also require a blank. This is a sample of the buffer that the DNA is in that allows the machine to establish background measures.
Once the blank and sample have been read by the machine the output will look something like this:
A260: 19.917 A280: 11.425 A260/A280: 1.74 A260/A230: 2.18
What does A260 tell me?
DNA absorbs light most strongly at 260 nm wavelength. So absorbance at 260 nm (or A260) can be used to directly estimate the concentration of DNA. That’s thanks to one of those old physical laws that are also equations.
The Beer-Lambert law says that absorbance is determined by three factors: (1) the length of the path the light takes through a substance in cm, (2) the concentration of the substance, and (3) an extinction coefficient specific to the substance in questions sometimes called the molecules’ molar absorptivity or in equation form. These relationships can be rearranged and written as:
Concentration* = (Absorbance / Length of Light Path ) x Molar Absorptivity
If you’ve just done a spec reading all of these are known! Absorbance is the A260 value, molar absorptivity is 50 (μg/mL)-1cm-1 for double-stranded DNA (33 (μg/mL)-1cm-1 for rarer single-stranded DNA samples and 40 (μg/mL)-1cm-1 for RNA), and length of the light path is a constant specific to the spectrometer model used. So a double-stranded DNA sample with A 260 read of 19.917 in a spec with a 1 cm path length will have a concentration of 998.5 μg/mL.
*If you diluted your sample either to bring it up to volume or because super concentrated DNA is hard to pipette or because you’re doing a dilution series don’t forget to then multiply this number by the dilution factor.
What does the A260/A230 ratio tell me?
Sometimes in the process of extracting and purifying extra contaminants get added to the mix. Luckily A230 absorbance reading can pick up many of the worst offenders including guanidinium salt (used in silica columns) and phenol (used in phenol/chloroform extractions). If either of these were used during extraction, it’s a good idea to look at the A260/A280 ratio. Like above you want the A260 number to be bigger. A rule of thumb is to aim for a ratio of 1.5 or more.
What does the A260/A280 ratio tell me?
In extractions and purifications, the goal is to separate DNA from other biological materials. Chief among these are proteins. Unfortunately, proteins are particularly good at evading removal. Enter an absorbance measure at 280 nm. While DNA absorbs light best at 260 nm most proteins absorb strongly at 280 nm so this ratio is a good general measure of DNA purity in a biological sample. You want the A260 read to be much higher than the A280 read. A good guide is to aim for a ratio number between 1.7 and 2.0.
What does A320 tell me?
This is a more general measure of the overall turbidity (cloudiness due to suspended matter) of the solution. High turbidity can drive up all the absorbance readings ( A260, A280, and A23). So take a glance at the A320 to gauge how much you can trust your other absorbance measures. If the number is high you may want to dilute the sample and take another read.
Some labs will also use the A320 reading in their DNA concentration calculations – replacing the A260 measurement with an absorbance number equal to A260 minus A320.
More Math Please!
Ever wonder how absorbance is calculated? When light is directed at a sample by a spectrometer it’s at a specific and measured radiant power (I0), After passing through the sample the light power is again measured (I1). The ratio of !1/1O is called transmittance (T). This number is converted to the more commonly used absorbance value using the equation A=Log(1/T.)
Online Calculators Please!
Now that you know what’s going on behind the scenes use these calculators to help you determine or double-check your sample’s DNA concentration. You just need to know the type of DNA in your sample, the A260 read, the path length of your spec, and whether or not the sample was diluted. Super handy but you’ll still need to use your brain and knowledge to judge the overall quality of your sample and the reliability of the concentration calculation.
– Omni Calculator: DNA Concentration
– AAT Bioquest DNA Concentration Calculator
Many of our most visited and liked posts relate to lab math so we’re expanding our repertoire. Have a numbers question that you want us to post about? Let us know!




1 comment
Comments are closed.