The basic unit of all living organisms, from bacteria to humans, is the cell. Contained within these cells is a molecule called deoxyribonucleic acid (or DNA). This amazing molecule contains the blueprint used to build an organism – our genetic makeup, or genotype, controls our phenotype (observable characteristics). DNA is built from nucleotides. Each nucleotide has three basic parts: a phosphate group, a deoxyribose sugar, a nitrogenous base – A, C, G, or T. The sugar of one nucleotide is covalent bonded to the phosphate group of its neighbor, forming long strands of DNA. Purified DNA used in molecular biology applications, including genetic engineering, transformation, restriction digest, PCR, and more. But how do we get it out of the cell? First, we need to break open the cell wall and membranes, the nuclear membrane, and then separate the DNA from all the other stuff that is present in the cytoplasm. We do this in a process called lysis. Once the cells are broken open, then we can use the chemical properties of DNA to separate it from the rest of the cellular components.
So, what do we need to extract DNA from cells?
- First, we need a sample. This can be anything that has DNA, from microbes to plants to human cells.
- Next, we need a DNA Extraction Buffer to help us lyse, or break open, the cell and nuclear membranes, releasing the DNA. Most DNA extraction buffers contain a molecules that stabilize the sample’s pH, a chelating agent to inactivate the enzymes that chew up DNA, a detergent that dissolves lipids, like the cell membrane, and salt, which disrupts the interactions between DNA and water.
- Then, we need ice cold isopropyl alcohol, which helps us force DNA out of the cell lysate. This results in long insoluble fibers of nucleic acid that we can either spool out using a glass rod, or a centrifuge to concentrate at the bottom of a tube.
- Finally, we need a way to physically break the cell walls and membranes. We can use a mortar and pestle to grind the cells, freeze/thaw to rupture cells by forming ice crystals, or a sonicator to use high frequency sound to break open the cells.
Now, we’re ready to get started. We’re going to put our sample to our tube and add our DNA extraction buffer. Next, we then crush the tissue by grinding it with our pestle, which physically crushes the cells and releases the DNA. The detergents in the buffer aid in dissolving the cell membranes, further assisting in the liberation of DNA from the cells.
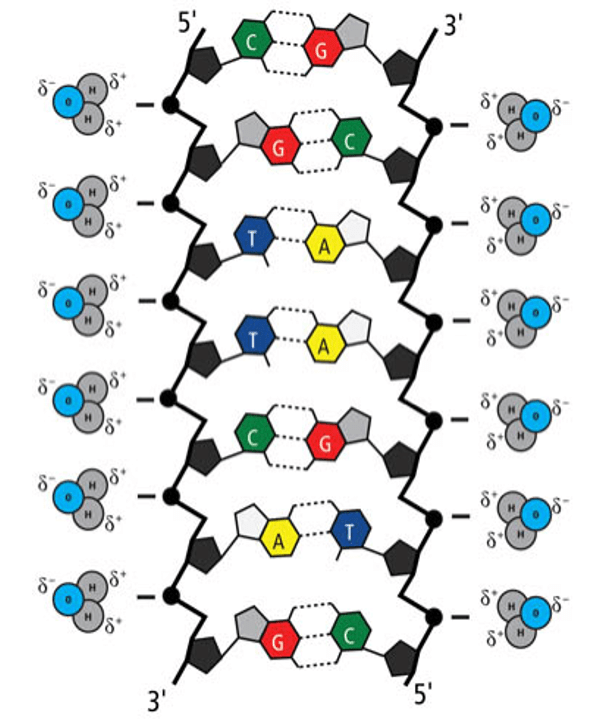
The salt, or sodium chloride, dissolved in our buffer exists as positively charged sodium ions and negatively charged chloride ions. The positive sodium ions neutralize the negative charge of DNA’s sugar-phosphate backbone. This improves its solubility in water. Without the salt, DNA remains negatively charged so it does not dissolve well in water.
Next, we add ice-cold isopropyl alcohol to the cell lysate, which causes the DNA to precipitate from cellular lysate as sticky white fibers due to its low solubility in isopropanol. The addition of salt neutralizes the negatively charged backbone of DNA, further decreasing its solubility. Furthermore, we’re going to make sure our alcohol is super cold, as the colder temperature promotes even greater DNA precipitation.
The precipitated DNA is separated from the rest of the cellular components in one of two ways. The first method involves layering the ice-cold isopropyl alcohol on top of the DNA solution, where the DNA will precipitate at the interface as sticky white fibers. To extract the DNA, a spooling rod is used to pull it out of the tube. The second and more common method is to mix and centrifuge the mixture at a very high speed, causing the purified DNA to form a pellet at the bottom of the tube. This pellet can be resuspended in buffer and used in molecular biology applications.
Here are some resources to help you in your DNA experiments!
- Conquering DNA Purity and Calculations
- Ten Things You Might Not Know About DNA
- Exploring our Chromosomal DNA
- Biotechnology Milestones – The Discovery of DNA



1 comment
Comments are closed.