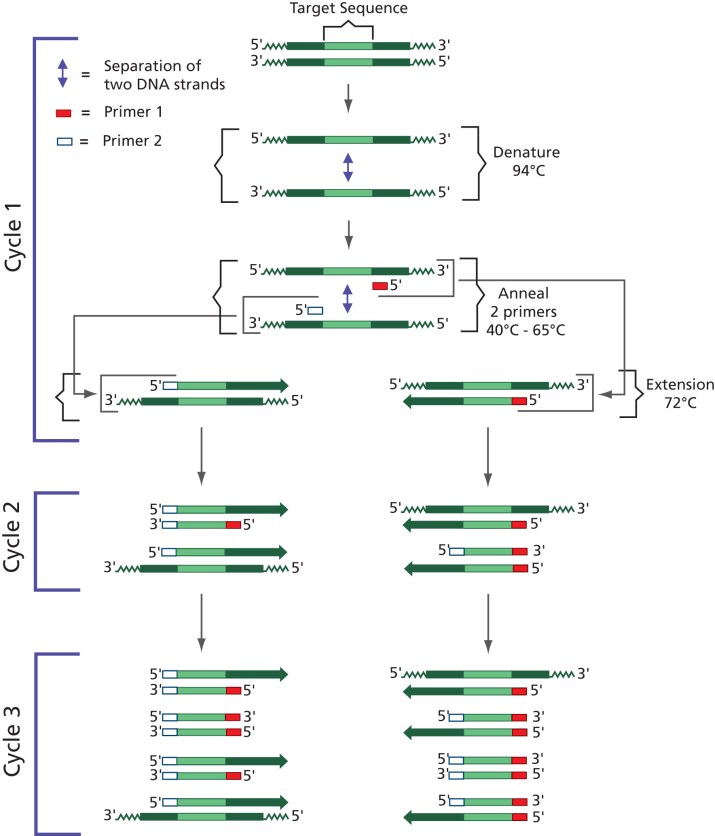
PCR (Polymerase Chain Reaction) is a widely used molecular biology technique that amplifies specific DNA segments. Researchers design primers that match the target DNA sequence, which is then amplified by Taq DNA polymerase. Basic PCR involves three main temperature steps – denaturation, annealing, and extension – that are repeated multiple times. This technique can amplify a DNA region as short as 70 base pairs or as long as a few kilobases.
Because of its ease of use and its ability to rapidly amplify DNA, PCR has become indispensable in the medical and life sciences lab. Over time, researchers began to adapt the technique to meet their needs. Below are a few of the most common adaptations of PCR technology.
- Reverse transcription PCR (RT-PCR): In the first step, the enzyme Reverse Transcriptase is used to convert RNA into complementary DNA (cDNA). The next step uses a basic PCR amplification protocol to target a sequence in the cDNA. RT-PCR is commonly used to study gene expression by measuring the amount of RNA present in a sample, or to diagnose viral diseases like HIV or COVID-19.
- Multiplex PCR: This technique allows the simultaneous amplification of multiple DNA targets in a single reaction. It is commonly used in diagnostic applications, such as detecting the presence of multiple pathogens in a single sample. We can perform multiplex PCR in the classroom, with experiments like our Water Quality Testing kit.
- Real-time or quantitative PCR (qPCR): This technique allows for the quantification of DNA in real time while the PCR sample is in the thermal cycler. It can be used for applications such as gene expression analysis and mutation detection. It is often paired with RT-PCR for applications like the detection of viral pathogens (this can be called qRT-PCR).
- Digital PCR: This technique splits a PCR sample into thousands of tiny droplets or chambers, allowing the quantification of very low levels of DNA or RNA. It can be used in applications such as rare mutation detection and quantification of viral load.
- Assembly PCR: This technique is used to assemble multiple DNA fragments into a single contiguous DNA molecule. It is commonly used in synthetic biology and genetic engineering applications.
- Nested PCR: This technique involves two rounds of PCR amplification. The first round amplifies a larger DNA fragment. This large fragment is then used as a template for the second round of PCR, which targets a smaller DNA sequence nested within the first.
- Inverse PCR: This technique is used to amplify unknown sequences that flank a known DNA segment. It is commonly used to study genomic rearrangements and identify the DNA sequences that are disrupted by the rearrangement.
These are some of the most commonly used types of PCR, but there are many other variations and modifications that have been developed to meet specific research needs.


1 comment
Comments are closed.