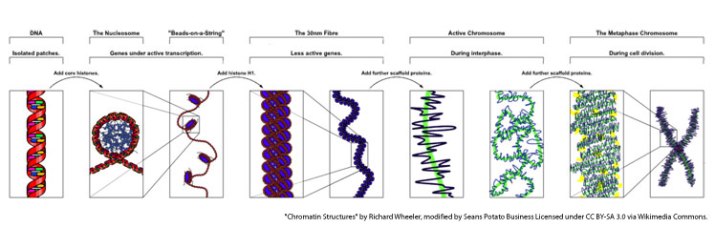
 The nucleus is a dense organelle that contains the genetic material of a cell. How dense? The human genome is around 3 billion nucleotides and the average length of a nucleotide is 0.6 nanometers, so stretched out the DNA in a cell would be around 1.8 meters or 5 ft. (Here’s the math: 3.0 × 109 x 6.0 × 10-10 meters = 1.8 meters). Cells must package 5 feet of genetic material into a nucleus of around 6 micrometers. They do this by tightly coiling the DNA around histone molecules to form chromatin. Chromatin is then coiled and looped and coiled and looped some more as illustrated above.
The nucleus is a dense organelle that contains the genetic material of a cell. How dense? The human genome is around 3 billion nucleotides and the average length of a nucleotide is 0.6 nanometers, so stretched out the DNA in a cell would be around 1.8 meters or 5 ft. (Here’s the math: 3.0 × 109 x 6.0 × 10-10 meters = 1.8 meters). Cells must package 5 feet of genetic material into a nucleus of around 6 micrometers. They do this by tightly coiling the DNA around histone molecules to form chromatin. Chromatin is then coiled and looped and coiled and looped some more as illustrated above.
How many loops? That’s one question that a group of scientists recently tackled with a landmark study that begins to map how the human genome folds itself inside the nucleus of a cell. The answer – around 10,000 – is much lower than had previously been predicted. The study also found that most loops are relatively small, less than 2 million base pairs. This work adds support and detail to the hypothesis that DNA loops play a crucial role in gene regulation. Loops often bring together distant enhancers and promoters that dictate gene expression. These are sometime referred to as secret switches. As loops form and un-form, different “secret switches” are turned on and off.
The 3D map of nuclear DNA folding and looping in on itself was created using some fundamental tools of the trade. First the cellular chromatin structure was treated with formaldehyde to preserve its 3D structure. Next, restriction enzymes cut the DNA into tiny pieces and biotin was added to mark the newly cut ends. Neighboring ends were then ligated together. These ligation junctions were then sequenced. For more on restriction enzymes check out this previous blog or one of our several kits that introduces students to this technology (for example, DNA Fingerprinting Using Restriction Enzymes).

1 comment
Comments are closed.